Introduction
The histone gene family. Definition of gene family. Pseudogenes. (p. 170-171)
The birth and death of genes
As genome evolve, new genes are born and old genes die. "Birth & death evolution" was mainly developed and promoted by Masatochi Nei beginning in the early 1970s. Many new genes arise by gene duplication but most of them become pseudogenes within a few million years. Some evolve new functions by subfunctionalizaton or neofunctionalization. (pp. 172-174) [On the evolution of duplicated genes: subfunctionalization vs neofunctionalization]
Box: The smell of sweat (pp. 174-175)
Gene duplication and mutationism
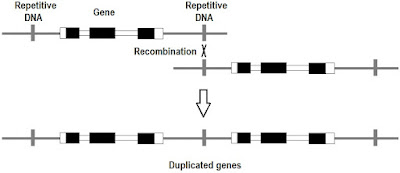
Gene duplication is due mostly to errors in recombination. This is a subset of segmental duplication and it leads to genome expansion. The creation of new genes by mutation is a key aspect of mutationism. (p. 175-177) [Mutation, Randomness, & Evolution] [Replaying life's tape] [What is "structuralism"?] [Reactionary fringe meets mutation-biased adaptation: Introduction]
Whole genome duplications and the fate of genes
Polyploidization and hybridization give rise to species with twice as much DNA. The fate of that extra DNA, especially extra genes, can be tracked over time. It looks like the extra DNA is another example of junk DNA, lending support to the idea that species can tolerate large amounts of nonfunctional DNA. (pp. 177-179) [The birth and death of salmon genes] [Birth and death of genes in a hybrid frog genome]
Box: Real orphans in the human genome
Completely new genes, de novo genes, are rare but there are genuine examples of genes that are unique in the human genome (ORFans). They arise by gene duplication and they are often polymophic. (p. 180)
Different kinds of pseudogenes
There are four different kinds of pseudogenes: death of a duplicated gene, processed, unitary, and polymorphic. The human genome has about 15,000 pseudogenes (5% of the genome) and almost all of them are junk. The fixation of a pseudogene involves two steps; mutation and fixation by random genetic drift. Pseudogenes can become unrecognizable after 100 million years. (pp. 181-184) [Is the high frequency of blood type O in native Americans due to random genetic drift?]
Box: Conserved pseudogenes and Ken Miller's argument against intelligent design
The presence of a conserved pseudogene in the beta globin gene cluster in chimpanzee and human genomes is difficult to explain by intelligent design. The fact that a small segment of the beta-globin pseudogene contains a SAR sequence is irrelevant to the main argument. (pp. 185-186)
Are they really pseudogenes?
Pseudogenes are broken genes and they are junk by any reasonable definition (see "If It Walks Like a Duck" in chapter 3). Some scientists who are opposed to junk DNA have claimed that most pseudogenes must be functional based on the fact that a tiny nunmber have secondarily acquired a functon. This is an example of cherry picking. (p. 186-188) [Are pseudogenes really pseudogenes?]
Box: The short legs of dachhunds (p. 188-189)
How accurate is the genome sequence?
The accuracy of DNA sequencing methods is approaching 99.99%. If that is coupled to 30x coverage, the overall accuracy is good enough to reliably distinguish between functional genes and pseudogenes. You also need a reliable sequence of your personal genome if you are going to make decisions about your health. (pp. 189-191)
Notes for Chapter 7 (pp. 327-328)