There are about 5000 noncoding genes. Some scientists attempt to solve The Deflated Ego Problem by postulating the existence of tens of thousands of new noncoding genes. (pp. 192-194) [John Mattick's new book]Different kinds of noncoding RNAs
In addition to tRNAs, rRNAs, and several unique RNAs, there are a number of classes of small RNAs: small nuclear RNAs (snRNAs); small nucleolar RNAs (snoRNAs); microRNAs (miRNAs); short interfering RNAs (siRNAs); PIWI-interacting RNAs (piRNAs); long noncoding RNAs (lncRNAs). (pp. 194-196)Understanding transcription
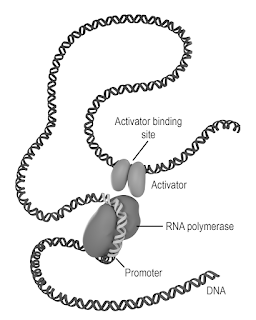
Transcription initiation requires the binding of RNA polymerase to a gene promoter. This is often aided by additional transcription factors that bind to nearby binding sites. (pp. 197-199)
Box: Three RNA polymerases for three different kinds of genes
On the important properties of DNA-binding proteinsTranscription factors bind to relatively small DNA sequences and these sequences will be present by chance in large genomes. Most transcription factors will be bound to nonfunctional sites. (pp. 200-203)Random transcription initiation is the rule
Eukaryotic cells will contain a lot of junk RNA produced by spurious transcription from nonfunctional sites. (pp. 203-204) [Sloppiness in translation initiation]Random transcription termination is also common
Transcription termination is inefficient leading to large amounts of junk RNA from the 3′ end of the gene. (p. 204)Sometimes RNA polymerase goes off in the wrong direction
Spurious transcription can also occur when transcription fires off in the wrong direction. (pp. 204-205)Antisense transcription
Most antisense transciption is spurious but there are a number of examples of regulatory antisense RNAs. Upstream antisense RNAs are sometimes called enhancer RNAs (eRNAs) and a small number of genes might be regulated by eRNAs. (pp. 205-206)How much of our genome is transcribed?
The ENCODE results from 2007 suggested that most of the human genome is transcribed at some time or other (pervasive transcription). It was assumed in 2007 that most of these transcripts were functional. (pp. 206-209) [A 2004 kerfuffle over pervasive transcription in the mouse genome] [Transcription activity in repeat regions of the human genome] [The pervasive transcription controversy: 2002]The pre-ENCODE history of pervasive transcription
The discovery that most of the human genome is transcribed dates back to the late 1960s. By 1980 it was understood that much of this is due to intron sequences that are rapidly degraded in the nucleus. There are two views on the possible function of all those transcripts; they are products of an exquisitely desiged genome like a Swiss watch or products of a sloppy genome. (pp. 209-211)How do we know about pervasive transcription?
DNA sequencing technology, RNA-Seq, and newer methods of detecting low level transcripts. (pp. 211-211) [The history of DNA sequencing]How many lncRNAs?
LncRNA databases contain more that 500,000 putative lncRNAs. Only a small number of lncRNAs have been shown to have a biologically relevant function. Scientists who claim that the genome is full of lncRNA genes often don't understand the concept of the null hypothesis. (pp. 212-217) [Most lncRNAs are junk] [Junk DNA [and lncRNAs]] [On the misrepresentation of facts about lncRNAs] [lncRNA nonsense from Los Alamos] [How many lncRNAs are functional?] [Experts meet to discuss non-coding RNAs - fail to answer the important question] [Confusion about the number of genes] [How many lncRNAs are functional: can sequence comparisons tell us the answer?]
Box: Revisiting the Central Dogma?
John Mattick proves his hypothesis?Many scientists are confused about the meaning of the central dogma of molecular biology and this confusion leads them to promulgate misconceptions about junk. (pp. 217-218) [Why is the Central Dogma so hard to understand?] [Subhash Lakhotia: The concept of 'junk DNA' becomes junk] [Georgi Marinov reviews two books on junk DNA]
John Mattick claims that the human genome contains huge numbers of regulatory noncoding genes demonstrating that the central dogma is wrong and junk DNA is a myth. The Human Genome Organizaton awarded him a major prize in 2012 for "proving" his hypothesis. Definition of "paradigm shaft." (pp. 218-221) [John Mattick presents his view of genomes] [The biggest mistake in the history of molecular biology (not!)] [John Mattick's latest attack on junk DNA] [Paradigm shifting]The null hypothesis
The importance of the null hypothesis (no function) and criteria for determining whether a transcript is functional of not. (pp. 221-224)
. Box: The Random Genome Project
On the origin of new genesNew genes (de novo genes) can arise from junk DNA and this process might be enhanced by the presence of spurious transcripts. This is a form of exaptation. Some scientists argue that the presence of excess DNA and pervasive transcription might be evolutionarily advantageous to a species. But there aren't very many examples of real de novo genes and the argument is teleological. (pp. 225-228) [Origin of de novo genes in humans] [The evolution of de novo genes] [Contingency, selection, and the long-term evolution experiment]
Box: Constructive neutral evolution (pp. 228-229)
What the scientific papers don't tell youMuch of the scientfic literature promotes the idea that there are tens of thousands of noncoding RNAs. They don't mention the fact that function as only been established for a small number of these transcripipts and they don't mention that spurious transcription is a reasonable explanation of pervasive transcription. (pp. 229-321)
Box: The false logic of the argument for noncoding RNAs (p. 232)
Biochemistry is messyBiochemical reactions are not perfect and errors are common. The cell does not look like a finely-tuned Swiss watch. (pp. 233-234)Change your worldview
If you think that every feature of a cell must be explained by adaptation then you should serously consider changing your worldview. Richard Dawkins and Stephen Jay Gould represent the two different views of evolution: adaptationism and pluralism. (pp. 234-237)Notes for Chapter 8 (pp. 328-330)
No comments:
Post a Comment